

 about
projects
people
publications
resources
resources
visit us
visit us
search
search
about
projects
people
publications
resources
resources
visit us
visit us
search
search
Quick Links
Featured Citations
Antiviral signaling of a type III CRISPR-associated deaminase. Li Y, Li Z et al. Science. 2025 Feb 21;387(6736):eadr0393.
The structure of apolipoprotein B100 from human low-density lipoprotein. Berndsen ZT, Cassidy CK. Nature. 2025 Feb 20;638(8051):836–843.
Bat genomes illuminate adaptations to viral tolerance and disease resistance. Morales AE, Dong Y et al. Nature. 2025 Feb 13;638(8050):449–458.
C-terminal amides mark proteins for degradation via SCF-FBXO31. Muhar MF, Farnung J et al. Nature. 2025 Feb 13;638(8050):519–527.
Resolving native GABAA receptor structures from the human brain. Zhou J, Noviello CM et al. Nature. 2025 Feb 13;638(8050):562–568.
More citations...News
December 25, 2024

|
December 12, 2024
The ChimeraX 1.9 production release is available! See the change log for what's new.
October 14, 2024
Planned downtime: The ChimeraX website, Toolshed, web services (Blast Protein, Modeller, ...) and cgl.ucsf.edu e-mail will be unavailable starting Monday, Oct 14 10 AM PDT, continuing throughout the week and potentially the weekend (Oct 14-20).
Previous news...Upcoming Events
UCSF ChimeraX (or simply ChimeraX) is the next-generation molecular visualization program from the Resource for Biocomputing, Visualization, and Informatics (RBVI), following UCSF Chimera. ChimeraX can be downloaded free of charge for academic, government, nonprofit, and personal use. Commercial users, please see ChimeraX commercial licensing.
ChimeraX is developed with support from National Institutes of Health R01-GM129325, Chan Zuckerberg Initiative grant EOSS4-0000000439, and the Office of Cyber Infrastructure and Computational Biology, National Institute of Allergy and Infectious Diseases.
Feature Highlight
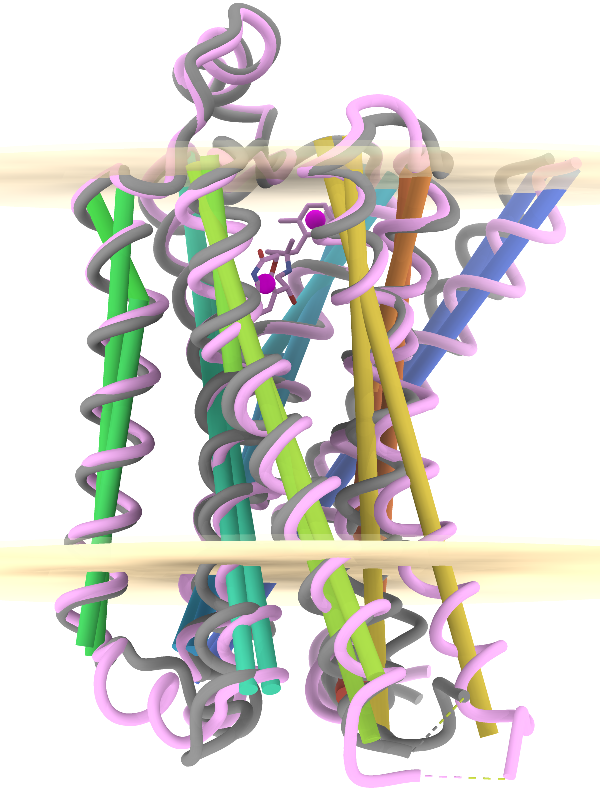
Axes, planes, and centroids can be defined from sets of atoms using the Axes/Planes/Centroids tool or the command define. The image shows the β2-adrenergic receptor in inactive (gray) and activated (pink) conformations, PDB 2rh1 and 4lde respectively. Hydrophobic membrane boundaries (light tan) are from the Orientations of Proteins in Membranes (OPM) database.
Plane objects were calculated from the membrane-boundary pseudoatoms in 4lde-OPM.pdb. Axes were calculated for the α-helices in both structures and rainbow-color-coded from blue at the N-terminus to red at the C-terminus. Centroids (magenta) were calculated from the two ring systems of the agonist in the activated structure. For image setup other than position, see the command file axes-gpcrs.cxc.
Distances and angles between the defined objects can be measured using the Axes/Planes/Centroids tool or commands, for example, to reveal that the ring systems of the agonist are about 9 Å apart and 3.3 and 8.7 Å from the outer membrane boundary; that upon activation, TM6 (gold axes) tilts by about 15° away from the helix bundle; and that TM4 (green axes, left side of image) makes an angle of about 80-85° with the membrane (see axes-measurements.cxc).
More features...Example Image
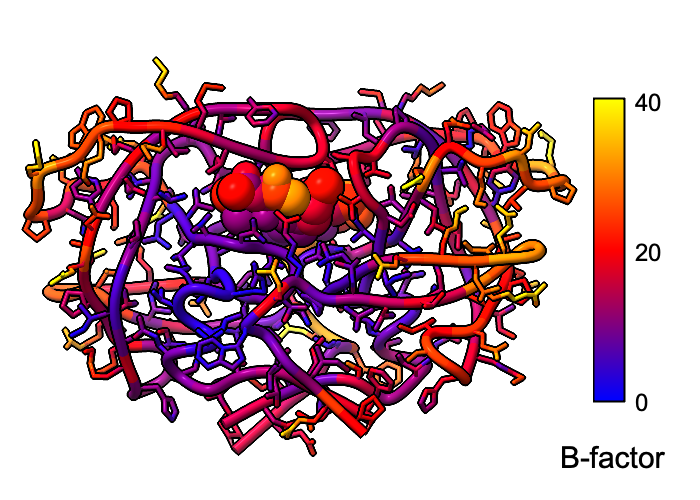
Atomic B-factor values are read from PDB and mmCIF input files and assigned as attributes that can be shown with coloring and used in atom specification. This example shows B-factor variation within a structure of the HIV-1 protease bound to an inhibitor (PDB 4hvp). For complete image setup, including positioning, color key, and label, see the command file bfactor.cxc.
Additional color key examples can be found in tutorials: Coloring by Electrostatic Potential, Coloring by Sequence Conservation
About RBVI | Projects | People | Publications | Resources | Visit Us
Copyright 2018 Regents of the University of California. All rights reserved.