 |
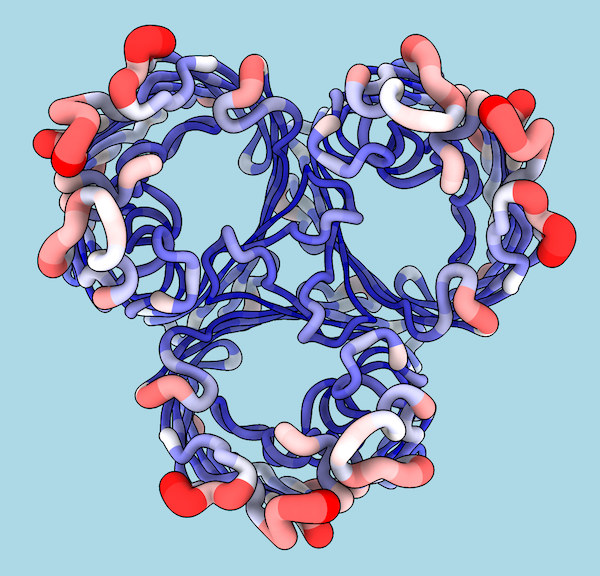
| 1pho coloring and worms by B-factor (more...) |
Render/Select by Attribute performs functions according to the values of attributes (built-in or user-defined):
Render/Select by Attribute is the graphical interface to the commands
color byattribute,
size byattribute,
cartoon byattribute,
and select (with command-line
specification by attribute).
See also:
Surface Color,
Color Key, the color bfactor icon
![]() in the Molecule Display Toolbar,
the Select menu,
measurements
in the Molecule Display Toolbar,
the Select menu,
measurements
Render/Select by Attribute set to its Render tab can be opened from the Depiction or Structure Analysis section of the Tools menu and manipulated like other panels (more...). Choosing Select... By Attribute Value... from the main ChimeraX menu opens the tool set to its Select tab.
Choosing Model and Attribute
Histogram or List of Attribute Values
Colors
Radii
Worms
The Models list includes all open atomic models. Individual models or blocks of models can be chosen with the left mouse button. Ctrl-click toggles the status of an individual model. To choose a block of models without dragging, click on the first (or last) and then Shift-click on the last (or first) in the desired block.
The Attribute of menu specifies whether the attribute of interest is at the level of atoms, residues, or structures (atomic models). The Attribute can then be chosen from the list of attributes available within the chosen models.
In the Render tab, only numerical attributes are available, whereas the Select tab also allows using boolean and string-valued attributes. These may include:
Custom attributes of atoms, residues, and molecule models can be created arbitrarily with defattr and setattr and (if numerical) mapped to colors with this tool.
In the Render section:
Choosing an attribute as described above shows a histogram of its values in the chosen models.Each threshold or vertical bar on the histogram has a Value (position on the histogram). It also has an associated Color (if the Colors tab is chosen), Atom radius (if the Radii tab is chosen), or Worm radius (if the Worms tab is chosen). Together, the thresholds specify how the colors or radii should map to values. Clicking on a threshold shows its Value in black. Clicking elsewhere within the histogram shows the Value (X-coordinate) of the mouseclick in gray. A threshold can be moved by changing its Value and then pressing Enter (return) or by dragging it horizontally with the left mouse button. Thresholds can be added by Shift-clicking on the histogram, whereas Shift-clicking an existing threshold deletes it.
Restrict to selection limits rendering by attribute to the current selection.
The Select tab of Render/Select by Attribute performs selection of atoms, residues, and structures (atomic models) based on their attribute values. In this section:
If the chosen attribute is numerical, the dialog shows a histogram of its values in the chosen models. The two thresholds or vertical bars on the histogram define a value range for selection, either between (inclusive) or outside their positions. Each threshold can be moved by dragging or directly entering a new value. The Value field applies to the threshold most recently moved or clicked. The no value option allows selecting items with no value assigned for the chosen attribute. For example, nucleic acid residues do not have peptide backbone phi angles.
If the chosen attribute is boolean, checkboxes allow specifying whether to select items for which the attribute is true or false.
If the chosen attribute is string-valued, a list of the values is shown. Individual rows or blocks of values can be chosen/unchosen by mouseclick or drag. Note that for the alt_loc attribute, only the values of currently used locations are listed, not all those present in the input file. Which alternate locations to use can be set with the AltLoc Explorer or the altlocs command.
Clicking Apply performs the action without closing the dialog, whereas OK performs the action and dismisses the dialog. Close simply dismisses the dialog, and Help shows this page in the Help Viewer.
The Value and Color settings apply to the threshold most recently moved or clicked. Clicking a color well opens the system color editor for choosing a color interactively. Coloring can be applied to atoms, cartoons, and/or molecular surfaces.
The Value and Atom radius settings apply to the threshold most recently moved or clicked. Rendering will change the VDW radii of the atoms. If needed, default VDW radii can be restored with the size command.
The Value and Worm radius settings apply to the threshold most recently moved or clicked. Attribute values are mapped to worm radius on a per-residue basis, and only biopolymer chains can be shown as worms. If an atom attribute is given, the average value over all atoms in a residue is used for that residue.
Once generated, the worm radii are remembered separately from the other cartoon parameters as a residue attribute named worm_radius. Clicking Deworm returns to the previous non-worm cartoon. The command cartoon byattribute (worm) can also be used to toggle back and forth between the previous non-worm cartoon and a previously defined worm display.